Many entity types can be uploaded in bulk using any common tabular format. Protein and nucleotide sequences can be imported using the GenBank format.
Upon upload, the registry will calculate and create any
molecular species and sets as appropriate.
Supported File Formats
The file formats supported are listed in the file import UI under the drop target area.
Tabular file formats are supported for all entity types:
- Excel: .xls, .xlsx
- Text: .csv, .tsv
Nucleotide sequences, constructs and vectors can also be imported using
GenBank file formats:
- GenBank: .genbank, .gb, .gbk
LabKey Biologics parses GenBank files for sequences and associated annotation features. When importing GenBank files, corresponding entities, such as new nucleotide and protein sequences, are added to the registry.
Assemble Bulk Data
When assembling your entity data into a tabular format, keep in mind that each Registry Source Type has a different set of required column headings.
Indicate Lineage Relationships
Lineage relationships (parentage) of entities can be included in bulk registration by adding "DataInputs/<DataClassType>" columns and providing parent IDs.
For example, to include a Vector as a 'parent' for a bulk registered set of Expression Systems, after obtaining the
template for Expression Systems, add a new column named "DataInputs/Vector" and provide the parent vector name for each row along with other fields defining the new Expression Systems.
Bulk Upload Registry Source Data
After you have assembled your information into a table, you can upload it to the registry:
- Go the Registry Source Type you wish to import.
- Select Add > Import from File.
- On the import page, you can download a template if you don't have one already, then populate it with your data.
- Confirm that the Registry Source Type you want is selected, then drag and drop your file into the target area and click Import.
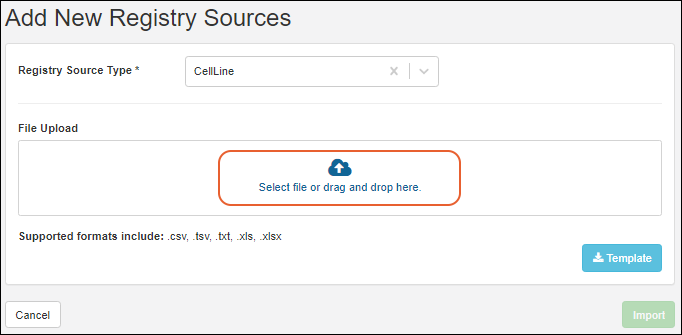
If you want to update existing registry sources or merge updates and creation of new sources, use
Edit > Update from File.
Bulk Data Example Files
Example Nucleotide Sequence File
Notes:
- Annotations: Add annotation data using a JSON snippet, format is shown below.
| name | alias | description | flag | protSequences | sequence | annotations |
|---|
| NS-23 | Signal Peptide 1 | An important sequence. | FALSE | [{name: "PS-23"}] | CCCCTCCTTG
GAGGCGCGCA
ATCATACAAC
CGGGCACATG
ATGCGTACGC
CCGTCCAGTA
CGCCCACCTC
CGCGGGCCCG
GTCCGAGAGC
TGGAAGGGCA | [
{
name:"First Annotation",
category:"Feature",
type:"Leader",
start:1,
end:20
},
{
name:"Another Annotation",
category:"Feature",
type:"Constant",
start:30,
end:50
}
] |
When importing the rows for NucSequence, you can reference the corresponding ProtSequence and the translation start, end, and offset. (The offsets are 1-based.) An example:
| name | alias | description | flag | protSequences | sequence | annotations |
|---|
| NS-100 | Signal Peptide 1 | some description | FALSE | [{name: "PS-100", nucleotideStart=1, nucleotideEnd=30, translationFrame=2}] | ATGGAGTTGGGACTGAGCTGGATTTTCCTTTTGGCTATTTTAAAAGGTGTCCAGTGT | |
Example Protein Sequence File
Notes:
- Organisms: A comma separated list of applicable organisms. The list, even if it has only one member, must be framed by square brackets. Examples: [human] OR [human, rat, mouse]
- ?: The column header for the extinction coefficient (ε).
- %?: The column header for the % extinction coefficient (%ε).
| Name | Alias | Description | Nuc Sequences | Chain Format | Avg. Mass | pI | ? | %? | Num. S-S | Num. Cys | Organisms | Sequence |
|---|
| PStest-150 | | Test sequence for import | | 1 | 13999.64 | 8.030 | 35500 | 2.54 | 1 | 2 | [mouse, rat] | EVQLVESGEL
IVISLIVESS
PSSLSGGLVQ
GGGSLRLSCA
ASGELIVISL
IVESSPSSLS
YSFTGHWMNW
VRQAPGKGLE
WVGIMIHPSD
SETRYNQKFK
DELIVISLIV
ESSPSSLSIR
FTISVDKSKN
TLYLQMNSLR
AEDTAVYYCA
RIGIYFYGTT
YFDYIWGQGT |
Mixtures and Batches
The text 'unknown' can entered for certain fields. For
Mixtures, the Amount field; for
Mixture Batches, the Amount and the RawMaterial fields.
Mixture Bulk Upload
| Type | Ingredient/Mixture | Amount Unit Type |
|---|
| Ingredient | I-2 | unknown |
Batch Bulk Upload
| Ingredient | Amount Used | Raw Material Used |
|---|
| Sodium phosphate dibasic anhydrous | 5 | RawMat-1234 |
| Sodium Chloride | unknown | unknown |
| Potassium chloride | unknown | unknown |
Downloadable Example Files
Download a sample file for a given entity type:
Related Topics
-
