When a Skyline document is imported, Panorama will parse and expose any annotation data, which can be used to create custom grids and visualizations.
Developers can determine where annotation data (such as Condition, BioReplicate, and Run) are stored using the
> Go To Module > Query. For example, peptide and small molecule annotations are stored in tables like targetedms.precursorannotation, targetedms.generalmoleculeannotation. All such tables have lookups (foreign keys) to the tables being annotated.
In addition, there is an "Annotations" column on each entity (targetedms.replicate, for example). In this column, all annotations are shown as name/value pairs. Upon import, any annotation columns will also be exposed as discrete "virtual" columns on the entity table itself. Such fields can then be exposed in grids using the UI, such as in the following example using replicate annotation data.
Replicate Annotation Data
Replicate annotation data is exposed in the query
targetedms.Replicate.
Users can access the annotation data by customizing their data grids:
- Starting from a data grid, select (Grid Views) > Customize Grid.
- In the Available Fields pane, open the node Sample File and then the node Replicate.
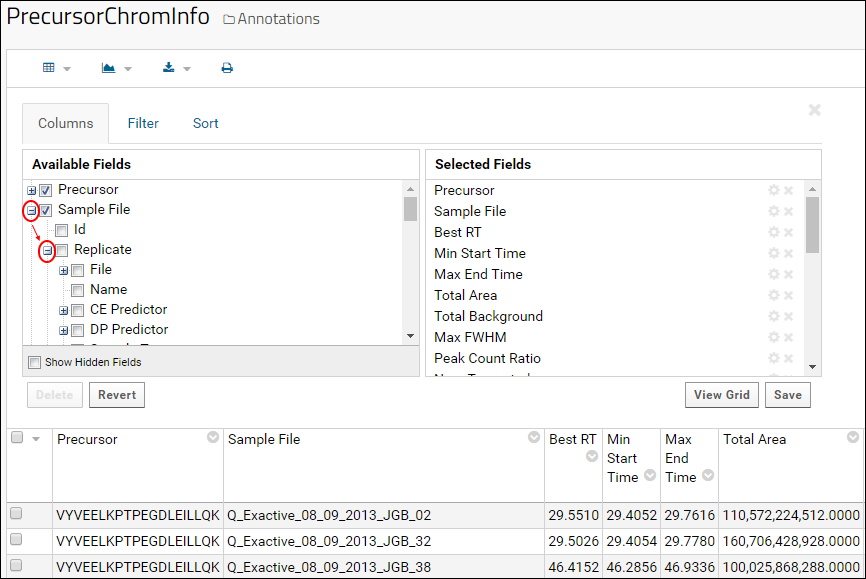
- Scroll down the Available Fields pane to see the available fields.
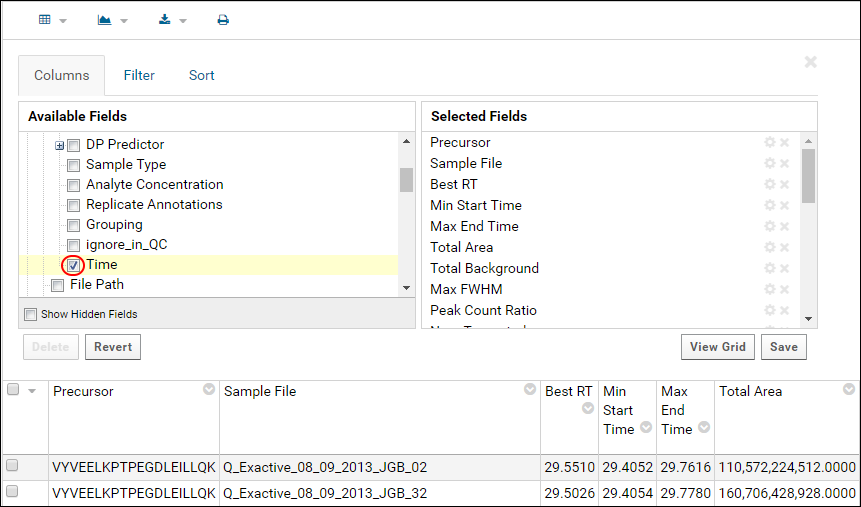
- Select fields to add them to the data grid. The screenshot below shows how to add the Time annotation field.